Tools such as RRDtool and rrd2csv can export an RRD file to xml or csv. While these formats can be imported into R, a more efficient and scalable approach would be to import the binary data directly (without exporting to an intermediate format first). To achieve this I implemented an R package that uses librrd to read binary data from an RRD file directly into data structures in R.
The .Call interface allows for calling C functions from R. Using .Call and the R headers for C makes it possible to pass R (SEXP) objects to C functions, generate, manipulate and return such back to R. A SEXP object can be of any data type used in R - e.g. vector, data.frame, integer, etc. This lets us implement C functions that take SEXP objects as arguments, use core librrd functions (rrd_fetch_r, rrd_info, rrd_first, rrd_last, etc.) to retrieve data from RRD files, generate SEXP objects (such as data.frame-s), populate them with the retrieved data and return them to the caller function in R.
Here is a great introduction to .Call written by Jonathan Callahan.
The package provides the following functions:
- importRRD(filename, consolidation function, start, end, step)
Acts as a wrapper around rrd_fetch_r. The user needs to have knowledge about the contents of the file beforehand to pick the right parameters. Returns a data.frame object containing the desired portion of the RRA that best matches the parameters. The data source names are retrieved and the columns of the data.frame are named accordingly. Due to the implementation specifics of rrd_fetch_r the result does not include the row with timestamp start.
For more information on rrd_fetch_r, please consult the official RRDtool documentation
- importRRD(filename)
Reads the metadata provided by rrd_info and uses it to import all RRA-s in their entirety. Retrieves an rrd_info_t struct which contains the parameters for each RRA and uses these, together with the boundary values provided by rrd_first and rrd_last to fetch each RRA using rrd_fetch_r. The user does not need to have any knowledge about the contents of the RRD file beforehand. Returns a list of data.frame objects, each labeled accordingly (“AVERAGE15” corresponds to an RRA with consolidation function “AVERAGE” and step 15).
- getVal(filename, consolidation function, step, timestamp)
When the user is interested in looking at individual values in an RRA it might not be convenient to retrieve the entire contents of an RRD if the file is too large. Retrieving portions of a specific RRA might result in a lot of reads from the RRD file (if few values are requested at a time) and indexing by row name in a data.frame (if more values are requested) is known to be inefficient in R.
getVal is optimized for working with indivudial values and uses a package-wide read-ahead cache to minimize the frequency of file reads. The cache is implemented as an environment object. Environments in R can be used as hash tables as is demonstrated here. A key is generated from filename, consolidation function and step. A data.frame is associated with each key and represents a per-RRA cache. The read-ahead size and the total size of the cache (per RRA) can be adjusted via setting the rrd.cacheBlock and rrd.cacheSize constants to the appropriate values.
Since looking up rows by row names (if we are to use dataFrame[“timestamp”, ]) is not done in constant time in R, the row index is calculated from the boundary timestamp values in the current cache and the timestamp and step parameters to allow for immediate indexing, instead. In order for this to work we have to make sure the cache contains no gaps - i.e. the time distance between any two adjacent values in the cache is step. getVal also checks if the timestamp is valid by looking at the boundaries of the cache and step. It will not cause an unnecessary call to rrd_fetch_r if it isn’t.
The cache is updated as the following:
– if rrd.cache[[key]] is NULL (i.e. there are no entires in the cache for that RRA) - retrieve a timestamp range that includes the rrd.cacheBlock next and previous values and store that in the cache.
– if the requested timestamp is larger than the latest currently stored timestamp in the cache, extend the cache to also include the values up until rrd.cacheBlock values after the requested one.
– similarly, extend the cache to include all values starting rrd.cacheBlock ones before the requested timestamp if it is smaller than the earliest currently stored.
– if the newly obtained cache entry for a given RRA is to exceed rra.cacheSize in size, that entry is replaced with the rrd.cacheBlock next and previous values around the requested timestamp as in the first case.
Another complication is due to the fact that, as mentioned above, rrd_fetch_r will try to always find the RRA that matches the parameters best. In case a value with a timestamp smaller than the earliest available for the RRA of interest is requested, rra_fetch_r will sometimes try to deliver a portion from another RRA in this RRD that contains a value with the specified timestamp. While this might be useful in certain situations, we really want to avoid this when working with the cache as each RRA we are getting values from is associated with a specific data.frame in the cache and we want all values to be step apart (the RRA that contains the timestamp could have a different step).
In order to solve this problem, getVal would obtain the earliest timestamp on the first cache-miss for a given RRA (and store it in a rrd.first environment object). It will then not allow rrd_fetch_r to be called with a timestamp smaller than the first one for a given RRA.
Example use
- Retrieving a portion from a certain RRA in a RRD file and plotting the data.
1 2 3 4 5 6 7 8 9 10 11 12 | |
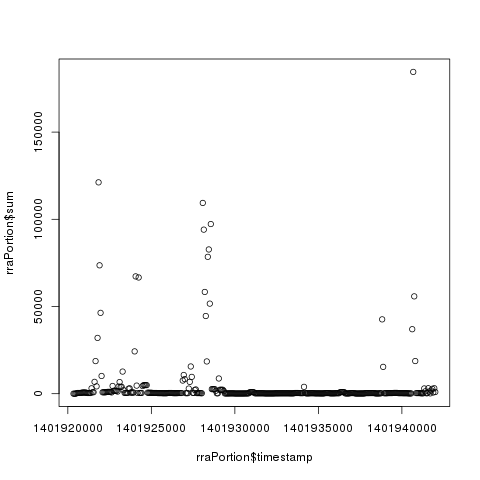
- Retrieving a selection of values at specific timestamps from a certain RRA.
1 2 3 4 5 6 7 8 9 10 11 12 13 | |
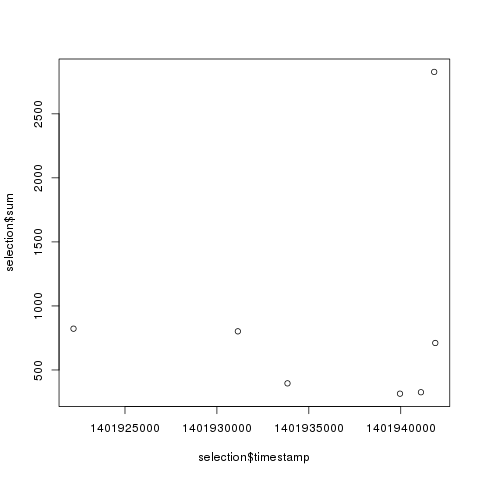
The same result can be achieved with:
1 2 3 | |
by importing the entire RRD file first.
- One can take advantage of the many possibilities to manipulate data.frame objects in R - e.g. performing various types of joins.
1 2 3 4 5 6 7 8 9 10 | |
This makes it possible to easily prepare metrics data from multiple data sources and RRD files to be used for statistical analysis or fed to a neural network.
The join_all function from the plyr package can be used to join an arbitrary number of data.frame objects
]]>